band_structure_of_nanocrystals
This is an old revision of the document!
Table of Contents
Band structure of nanocrystals
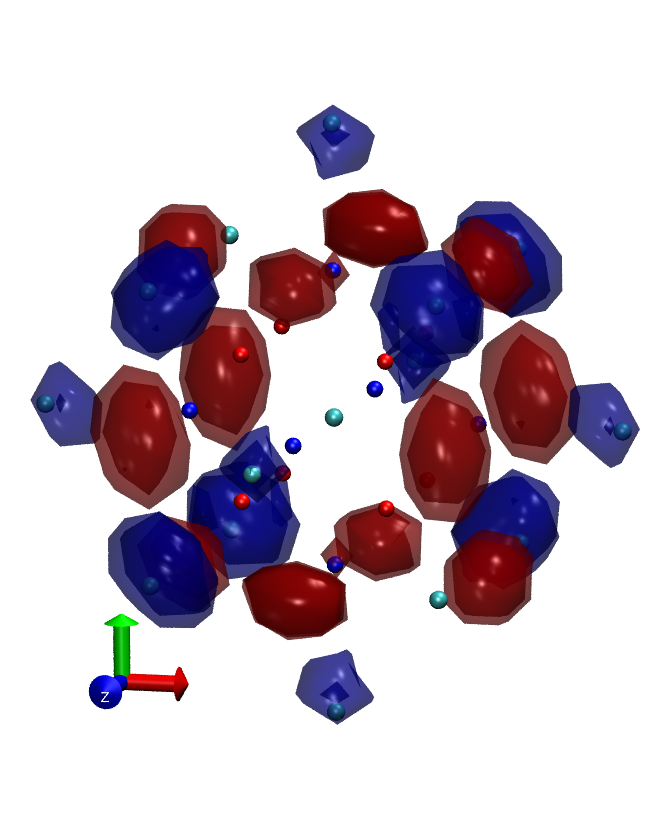
The 'fuzzy band structure' (i.e. cluster without periodic boundary condition) of an nanocrystal can be obtained by Fourier transform of it's eigenstates. See Hapala et.al,Phys.Rev.B87,195420. In fireball there are two methods to do this projection.
- 3D Fourier transform of molecular orbitals - in this method we choose several molecular orbitals, project them on grid and let fireball execute Fast fourier transform on this grid. The problem is that the result is complex ( real part correspond to symmetric components of the real wavefunction and imaginary part to antisymetric componenets ). For easier visualization we compute square of absolute value, which corresponds to density of the state in k-space. This property is exported as .xsf file.
Availability
this is implemented only in Prokop's personal versions of Fireball located in
/data/home/hapala/Fireball_dev/src_1.0-BSfinal /data/home/hapala/Fireball_dev/progs_Jellium_mod
3D Fourier fransform of Molecular Orbitals
fireball.in
&OPTION basisfile = answer.bas nstepf = 1 icluster = 1 ifixcharge = 1 dt = 0.5 &END &OUTPUT iwrtewf = 1 &END &MESH iewform = 5 npbands = 5 pbands = 159,160,161,162,163 &END
kmap.optional
1 # byEnerg - if 1 choose orbitals by energy range else choose by pbands list -8.59705 1.09186 # Emin Emax - energy range 0 # PlotXSF? - save .xsf files 0 # PlotPPN? - save 2D projectins (max along z-axis) into ppn image file for quick overview 0 # PlotImag? - save real and imaginery part to independent files fft_Im_XXXX.xsf and fft_Re_XXXX.xsf 1 # GetMaxPos - if 1 for each MO writes out position of global maximum of k-space density int kmaxs.dat 5.50 # alat - lattice constant in angstroem 0 # ValIn - write out value in different points defined in ksamples.optional
ksample.optional
26 nksamples 2.0 0.0 0.0 # HOMOs Gamma 100 0.0 2.0 0.0 0.0 0.0 2.0 -2.0 0.0 0.0 0.0 -2.0 0.0 0.0 0.0 -2.0 1.0 1.0 1.0 # HOMOs Gamma 111 -1.0 1.0 1.0 1.0 -1.0 1.0 -1.0 -1.0 1.0 1.0 1.0 -1.0 -1.0 1.0 -1.0 1.0 -1.0 -1.0 -1.0 -1.0 -1.0 0.0 1.0 1.0 # LUMOs X 0.0 -1.0 1.0 0.0 1.0 -1.0 0.0 -1.0 -1.0 1.0 0.0 1.0 -1.0 0.0 1.0 1.0 0.0 -1.0 -1.0 0.0 -1.0 1.0 1.0 0.0 -1.0 1.0 0.0 1.0 -1.0 0.0 -1.0 -1.0 0.0
kmaxs.dat
iband, eigen_k(iband,1), imax, jmax, kmax, ValMax, rimax, rjmax, rkmax 107 -7.40951516 -3 -3 -3 2968.71621697 -0.70854607 -0.70769619 -0.70733483 108 -7.38731466 -4 -3 -2 3133.46757266 -0.94472810 -0.70769619 -0.47155655 109 -7.38677828 -4 -2 -3 3128.34104148 -0.94472810 -0.47179746 -0.70733483 110 -7.26838823 -4 -4 0 1878.06049291 -0.94472810 -0.94359492 0.00000000 ...
ksamples.dat
Ei rho(k1) rho(k2) rho(k3) rho(k4) rho(k5) rho(k6) -2.08815 16.68657904 3.16266264 10.09450414 16.68657904 3.16266264 10.09450414 ... -2.08681 5.87885299 17.02497532 10.22090870 5.87885299 17.02497532 10.22090870 ... -2.06449 28.89941918 27.76432636 29.01794378 28.89941918 27.76432636 29.01794378 ... ...
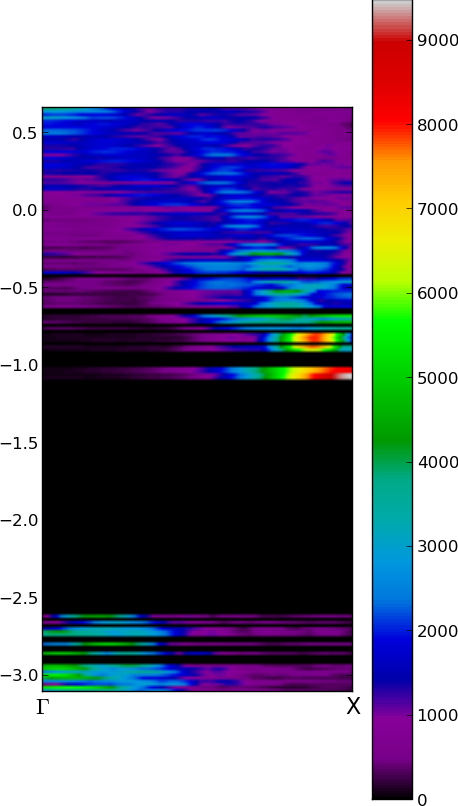
1D cuts of Fuzzy Band Structure
fireball.in
&OPTION basisfile = answer.bas nstepf = 1 icluster = 1 ifixcharge = 1 dt = 0.5 &END &OUTPUT iwrtewf = 1 &END &MESH iewform = 6 &END
kscan.optional
1 # byEnerg -8.0000 1.0000 # Emin Emax 5.50 # alat 48 100 # nlines nkpoints 2.0 0.0 0.0 1.0 1.0 0.0 2.0 0.0 0.0 1.0 -1.0 0.0 2.0 0.0 0.0 1.0 0.0 1.0 2.0 0.0 0.0 1.0 0.0 -1.0 0.0 2.0 0.0 1.0 1.0 0.0 0.0 2.0 0.0 -1.0 1.0 0.0 0.0 2.0 0.0 0.0 1.0 1.0 0.0 2.0 0.0 0.0 1.0 -1.0 0.0 0.0 2.0 -1.0 0.0 1.0 0.0 0.0 2.0 1.0 0.0 1.0 0.0 0.0 2.0 0.0 1.0 1.0 0.0 0.0 2.0 0.0 -1.0 1.0 -2.0 0.0 0.0 -1.0 1.0 0.0 -2.0 0.0 0.0 -1.0 -1.0 0.0 -2.0 0.0 0.0 -1.0 0.0 1.0 -2.0 0.0 0.0 -1.0 0.0 -1.0 0.0 -2.0 0.0 1.0 -1.0 0.0 0.0 -2.0 0.0 -1.0 -1.0 0.0 0.0 -2.0 0.0 0.0 -1.0 1.0 0.0 -2.0 0.0 0.0 -1.0 -1.0 0.0 0.0 -2.0 1.0 0.0 -1.0 0.0 0.0 -2.0 -1.0 0.0 -1.0 0.0 0.0 -2.0 0.0 1.0 -1.0 0.0 0.0 -2.0 0.0 -1.0 -1.0 1.0 1.0 1.0 0.0 1.0 1.0 1.0 1.0 1.0 1.0 0.0 1.0 1.0 1.0 1.0 1.0 1.0 0.0 -1.0 1.0 1.0 0.0 1.0 1.0 -1.0 1.0 1.0 -1.0 0.0 1.0 -1.0 1.0 1.0 -1.0 1.0 0.0 1.0 -1.0 1.0 0.0 -1.0 1.0 1.0 -1.0 1.0 1.0 0.0 1.0 1.0 -1.0 1.0 1.0 -1.0 0.0 -1.0 -1.0 1.0 0.0 -1.0 1.0 -1.0 -1.0 1.0 -1.0 0.0 1.0 -1.0 -1.0 1.0 -1.0 -1.0 0.0 1.0 1.0 -1.0 0.0 1.0 -1.0 1.0 1.0 -1.0 1.0 0.0 -1.0 1.0 1.0 -1.0 1.0 1.0 0.0 -1.0 1.0 -1.0 0.0 1.0 -1.0 -1.0 1.0 -1.0 -1.0 0.0 -1.0 -1.0 1.0 -1.0 -1.0 1.0 0.0 1.0 -1.0 -1.0 0.0 -1.0 -1.0 1.0 -1.0 -1.0 1.0 0.0 -1.0 1.0 -1.0 -1.0 1.0 -1.0 0.0 -1.0 -1.0 -1.0 0.0 -1.0 -1.0 -1.0 -1.0 -1.0 -1.0 0.0 -1.0 -1.0 -1.0 -1.0 -1.0 -1.0 0.0
klines_XXXXX.dat
1 391.30476603 391.30476603 391.30476603 391.30476603 287.45162513 ...
2 391.20959758 391.21921233 390.08793464 391.61800721 295.99428664 ...
3 389.17880205 389.77130848 387.81208053 388.10778349 306.53352273 ...
...
klines_TOT.dat
Ei rho(k1) rho(k2) rho(k3) rho(k4) rho(k5) ... -7.99755 413.91048549 428.76133304 440.87314746 450.14882297 456.52837031 ... -7.99140 394.18797633 403.02567283 409.86012222 414.38007661 416.29758353 ... -7.97472 418.51477832 433.83793597 448.50901654 462.18377663 474.53904963 ... -7.96729 262.72449682 269.69434762 274.26597685 275.89514172 285.23659456 ... ...
plotting results
#!/usr/bin/python
from pylab import *
import numpy
dE = 0.02 # [ eV ]choose width of energy-bins for plotting
F = transpose(genfromtxt( 'klines_TOT.dat' ))
Es = F[0]
Ps = transpose(F[1:])
Emin =Es.min(); Emax =Es.max()
Pgrid = zeros( ( int( (Emax-Emin)/dE )+1 , shape(Ps)[1] ) )
n = len(Es)
for i in range(n):
iE = int((Es[i]-Emin)/dE)
#Pgrid[iE] += Ps[i] # overlaping (=degenerated) states are summed up
Pgrid[iE] = numpy.vstack([Pgrid[iE],Ps[i] ]).max(axis=0) # for overlaping (=degenerated) states is taken maximum
# choose color map reference here http://wiki.scipy.org/Cookbook/Matplotlib/Show_colormaps
cmap='spectral'
figure( figsize=(5,10) )
extent = ( 0,2, Emin, Emax )
imshow( Pgrid, origin='image', extent=extent, cmap = cmap )
xticks([0,2] , ['$\Gamma$','X'], fontsize=16) # x-axis ticks Gamma and X vector
colorbar()
savefig('FuzzyBand.png', bbox_inches='tight' )
show()
band_structure_of_nanocrystals.1416317631.txt.gz · Last modified: 2014/11/18 14:33 (external edit)