This is an old revision of the document!
Table of Contents
Introduction
A Python/C based package, available at http://github.com/ProkopHapala/ProbeParticleModel/tree/PPSTM, which primary purpose is to simulate STM or dI/dV signal obtained with tilting tip apex (like CO, or Xe tip). It is based on the original PPAFM Python/C by Prokop Hapala and Co. http://github.com/ProkopHapala/ProbeParticleModel/ for simulation of tilting tip apex' AFM images.
Basic Principles
The PP-STM code calculates an STM or dI/dV signal based on the input from various Linear Combination of Atomic Orbitals (LCAO)-DFT codes. These inputs are eigen-energies of eigen-states (molecular orbitals) and LCAO coefficients (for each state and atomic orbital of sample). Sample can be approximated by freestanding molecule, or by full system - molecule/substrate.
Inputs for DFT Codes
At the moment the PP-STM code can read inputs from following DFT codes:
Fireball
A working executable for creation of necessary file is at: /storage/praha1/home/krejcio/bin_fireball_stable/fireball.x
Input files can computed with McWEDA functional as well as with computing XC on a grid. Cluster systems as well as systems with Periodic Boundary Conditions (PBC) can be computed. A fireball.in file for calculations with McWEDA:
&OPTION basisfile = 'answer.bas' lvsfile = 'input.lvs' kptpreference = 'input.kpts' ! only gamma point-calculations work at the moment nstepf = 1 icluster = 0 ! 0 for PBC / 1 for cluster calculation itdse = 0 iqout = 1 ifixcharge = 1 ! 0 if you don't have pre-calculated atomic charges in CHARGES iquench = -1 &END &OUTPUT iwrtcdcoefs = -2 ! print the important files &END
A fireball.in file for calculations with XC on a grid computations:
&OPTION
basisfile = 'answer.bas'
lvsfile = 'input.lvs'
kptpreference = 'samplek.kpts'
nstepf = 1
icluster = 0 ! 0 for PBC / 1 for cluster calculation
itdse = 0
iqout = 1
ifixcharge = 0
dt = 0.5
iquench = -1
iks = 1
imcweda = 0
idogs = 0
bmix = 0.05
&END
&OUTPUT
iwrtcdcoefs = -2
&END
! Next part is more important for calculating fftpot.xsf, which is Hartree potential important for PP-AFM calculations with charged tip apex.
&MESH
ifixg0 = 1 !
g0 = 0.0,0.0,0.0 ! do not shift the position of atoms in fftpot.xsf with respect to the answer.bas
Ecut = 300.0d0 ! not really necessary, but gives grid sampling approximately 100 pm.
&END
In case of PBC calculations phik_0001_s.dat, phik_0001_py.dat, … files are produced by the Fireball. In case of cluster calculations phik_s.dat, phik_py.dat, … are outputs of the Fireball calculations. They serve as inputs for the PP-STM calculations. Inside they look like:
38 280 -5.37896401 Number of atoms Number of states (Molecular orbitals) The Fermi Level -27.58251 -0.00004 0.00000 -0.00006 0.00000 0.00002 0.00000 ... Eigen-energy of the 1st state Real & Imaginary part for the LCAO coeficient for 1st state 1st atom for (s, py ... depending on the name of file) etc. -27.58139 0.02716 0.00000 0.04766 0.00000 0.00660 0.00000 ... eigen-energy of the 2nd state Real & Imaginary part for the LCAO coeficient for 2nd state 1st atom for (s, py ... depending on the name of file) etc. ...
GPAW
http://wiki.fysik.dtu.dk/gpaw/
Even though the GPAW is mainly used for representing the wave-function on a grid it can work in LCAO mode as well. For the purpose of making inputs for the PP-STM calculations the LCAO mode is necessary. Both - default or double-zeta (basis='dzp'; for more information look at the GPAW web page) - basis sets can be used. The PP-STM code reads the stored *.gpw binary produced by the GPAW calculations. Here is an example of some GPAW script for the calculations of the input:
from ase import *
from ase.visualize import *
from ase.io import*
from gpaw import *
import numpy as npy
mol = read('input.xyz') # xyz geometry of the sample
cell = npy.loadtxt('input.lvs') # cell in which a sample is
mol.set_cell(cell)
mol.set_pbc(False) # cluster calculation, but PBC can be used as well: mol.set_cell(cell)
mol.center()
xc='LDA' # other XC like PBE, RPBE, PW91, BLYP can be used, too.
calc = GPAW(txt='out_LCAO.txt',xc=xc,mode='lcao',basis='dzp')
mol.set_calculator(calc)
en = mol.get_potential_energy()
print en
calc.write('out_LCAO_'+xc+'.gpw',mode='all') # saves the calculation into binary 'out_LCAO_LDA.gpw' file
The results of the GPAW calculations is stored in 'out_LCAO_LDA.gpw'
FHI-AIMS
http://aimsclub.fhi-berlin.mpg.de/
Works for PBC calculations, just add:
output eigenvectors output band 0 0 0 0.5 0.5 0.0 3 G K
into control.in.
Note: For cluster calculations Mathematica scripts have to be used for creating PP-STM inputs.
Example
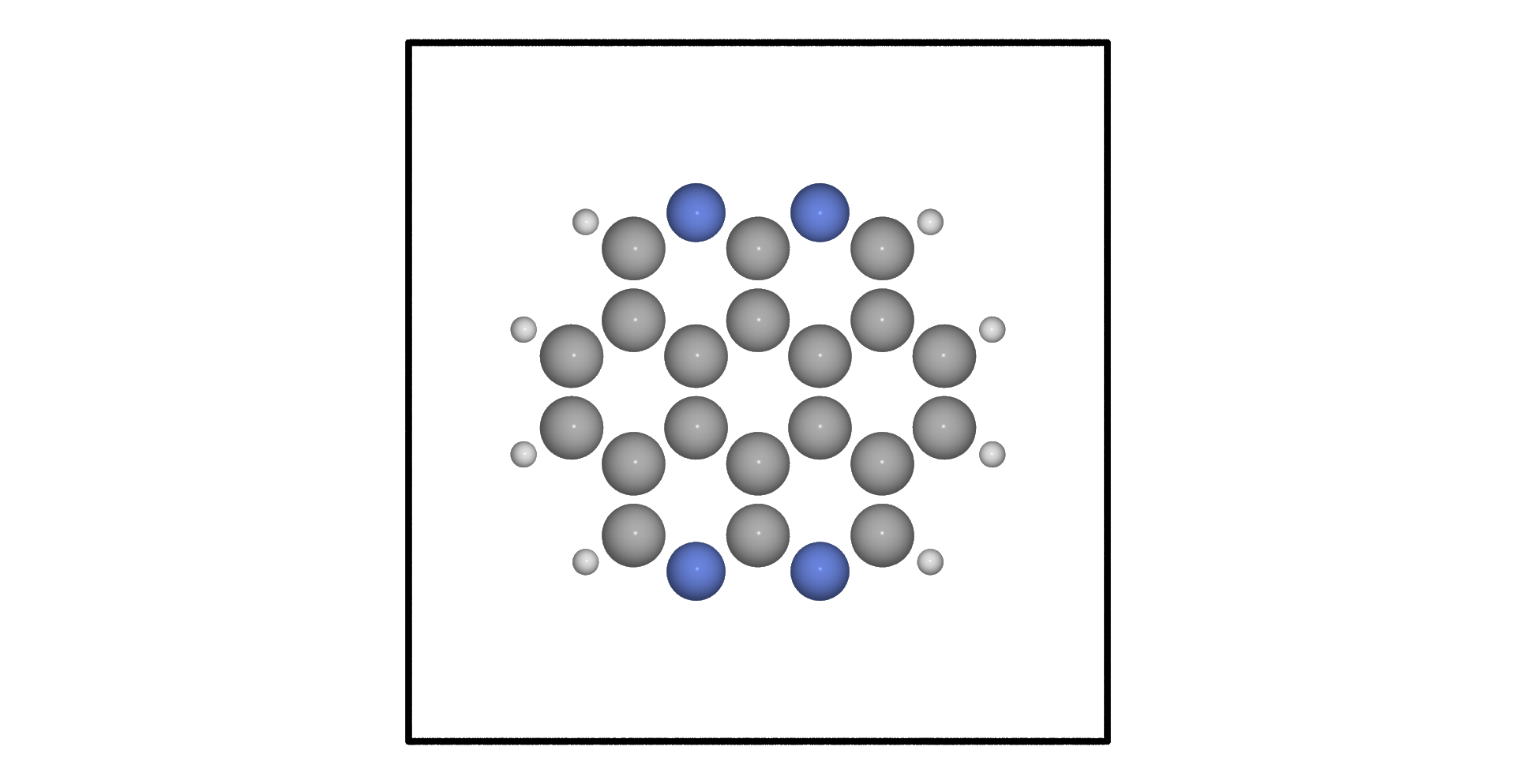
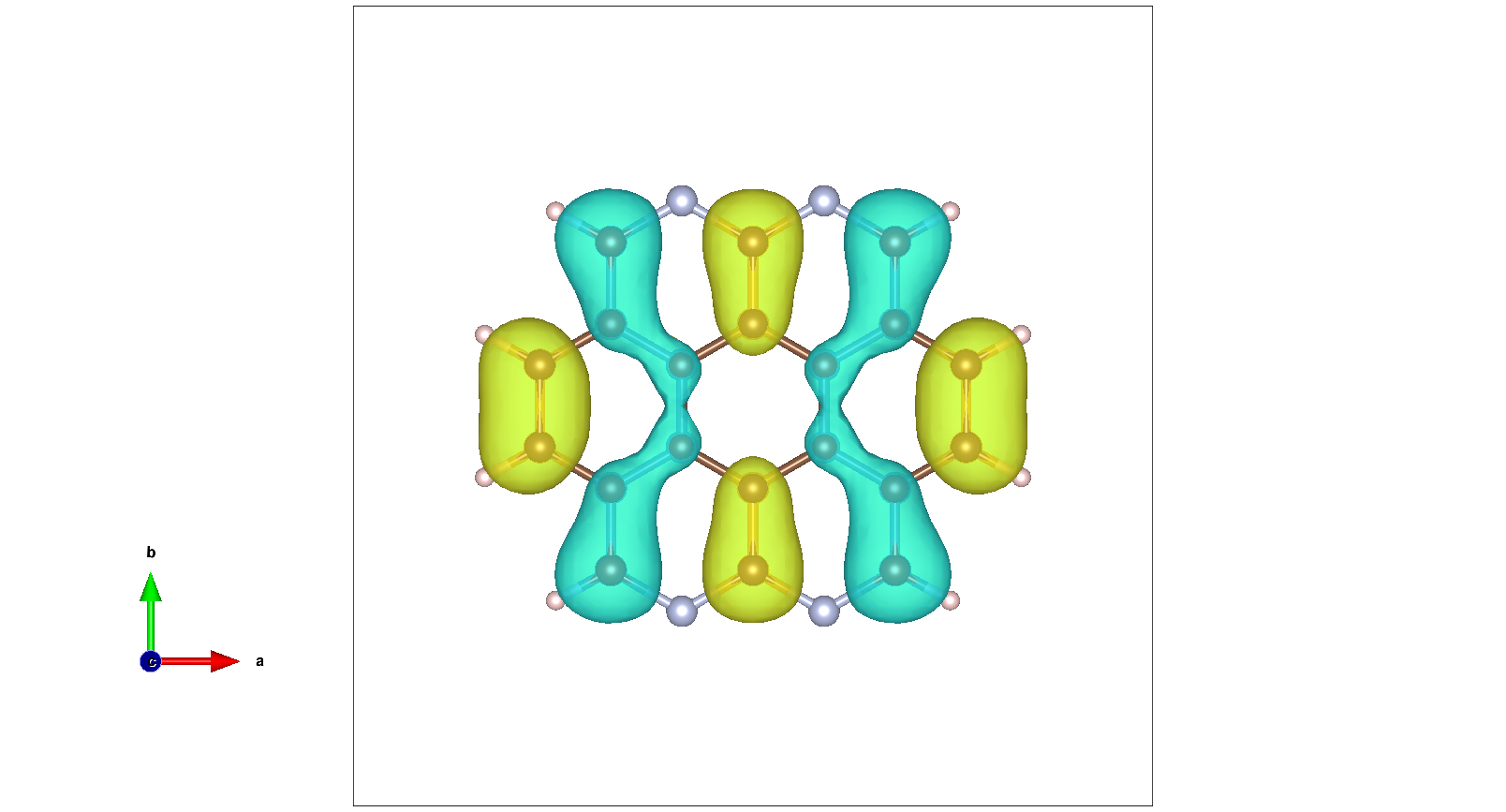
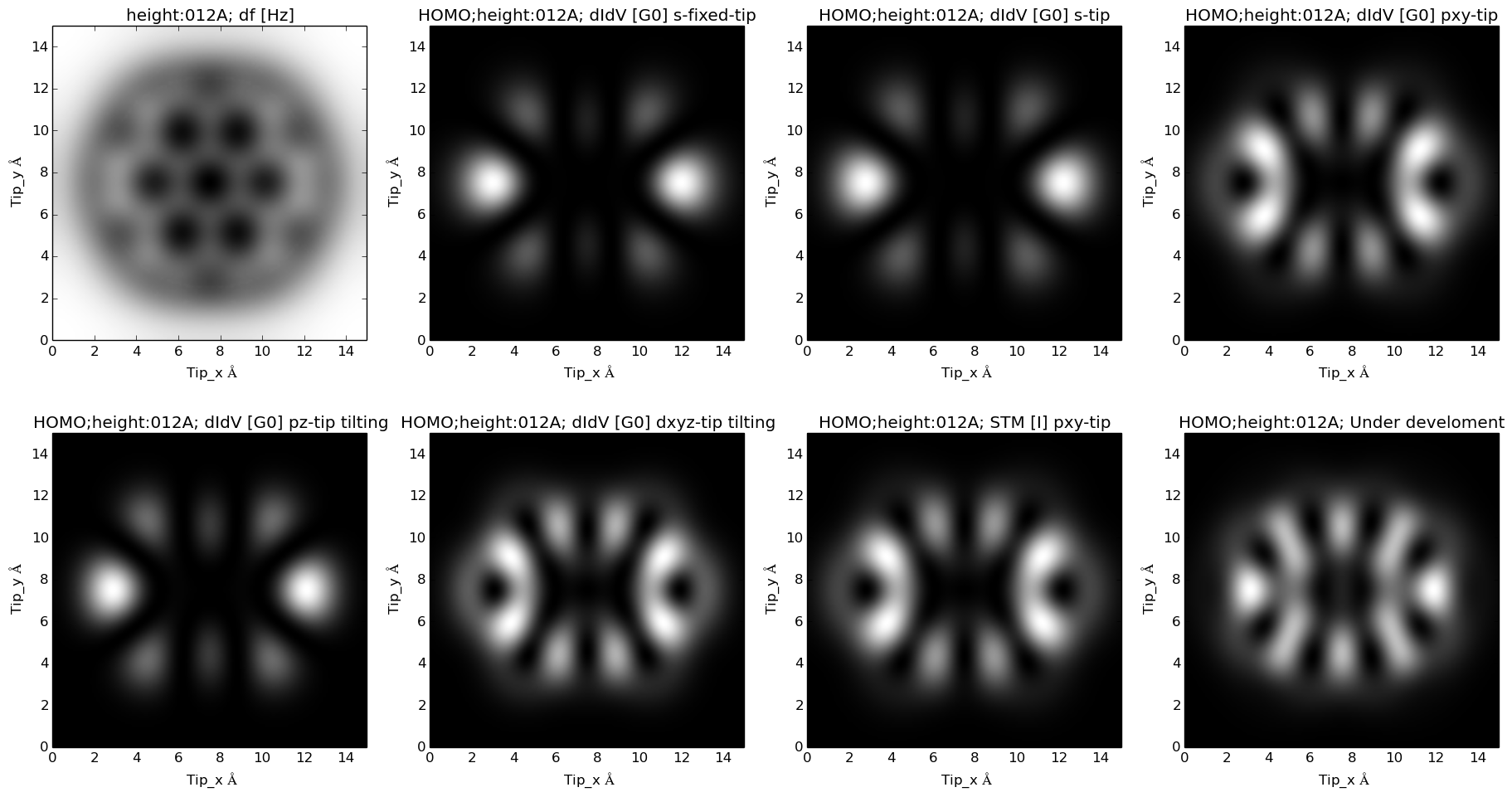
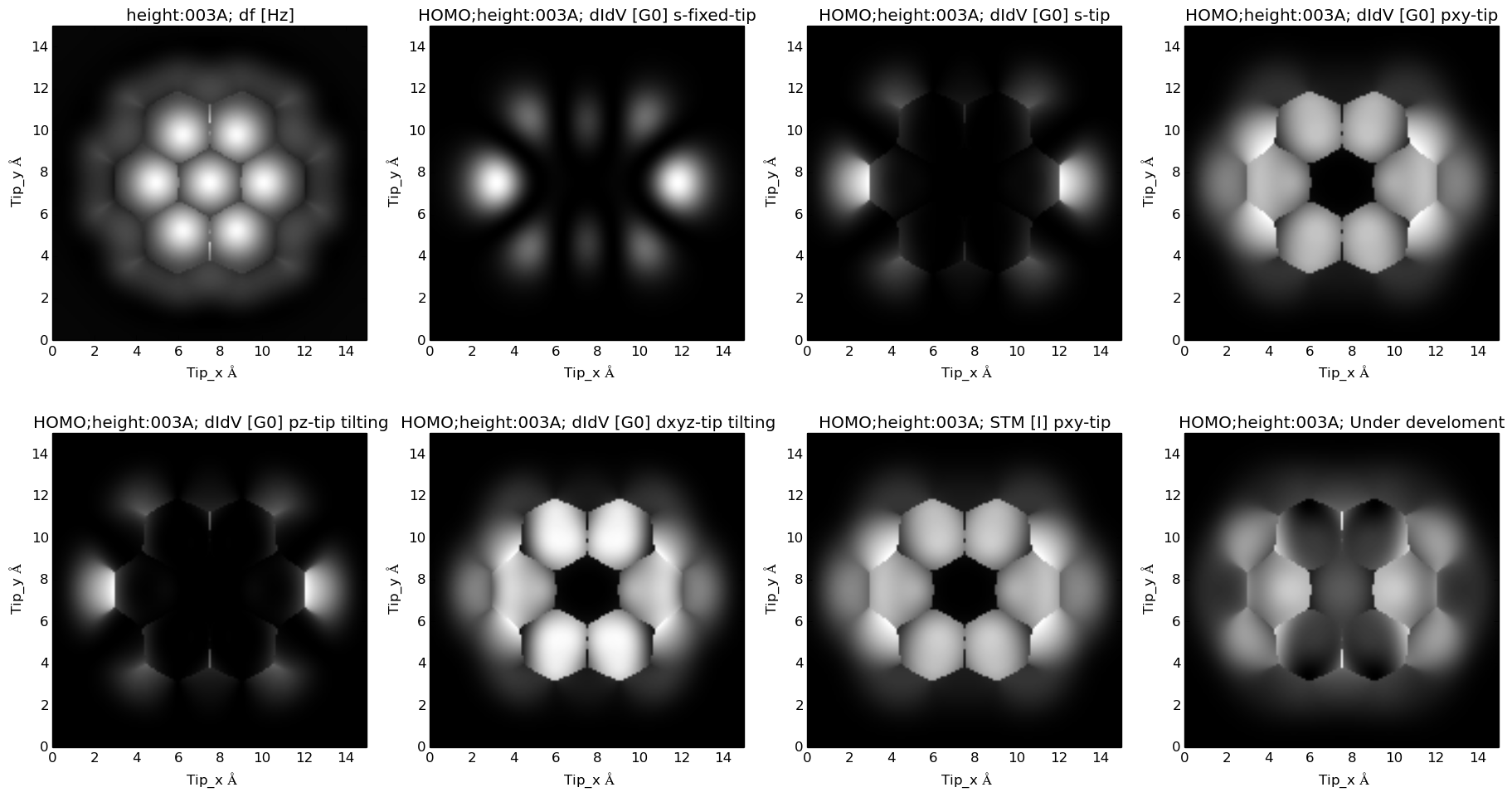
Theoretical prediction for Coronene molecule modified with with four nitrogen atoms :
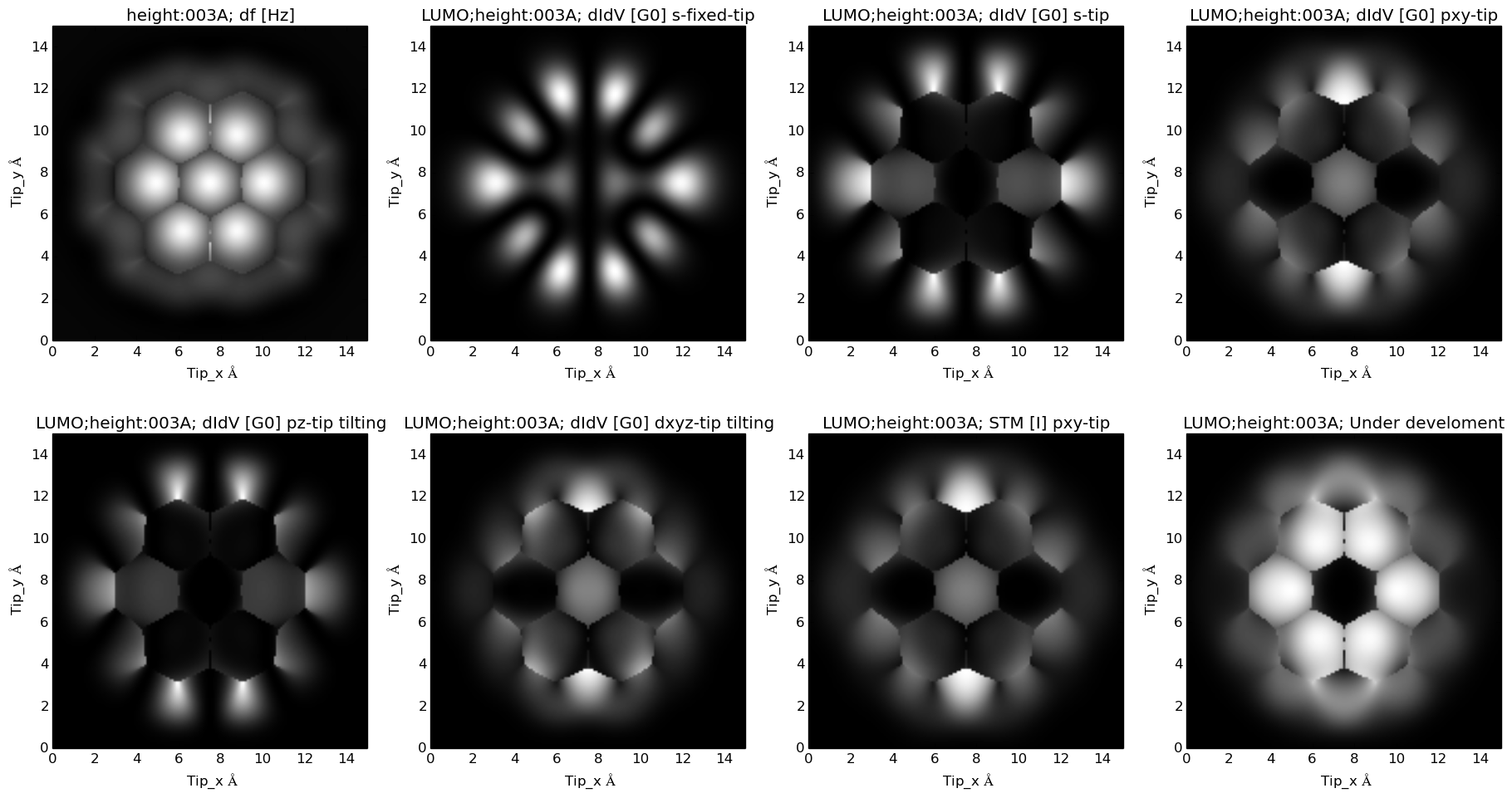
LDA predicted HOMO for this molecule:
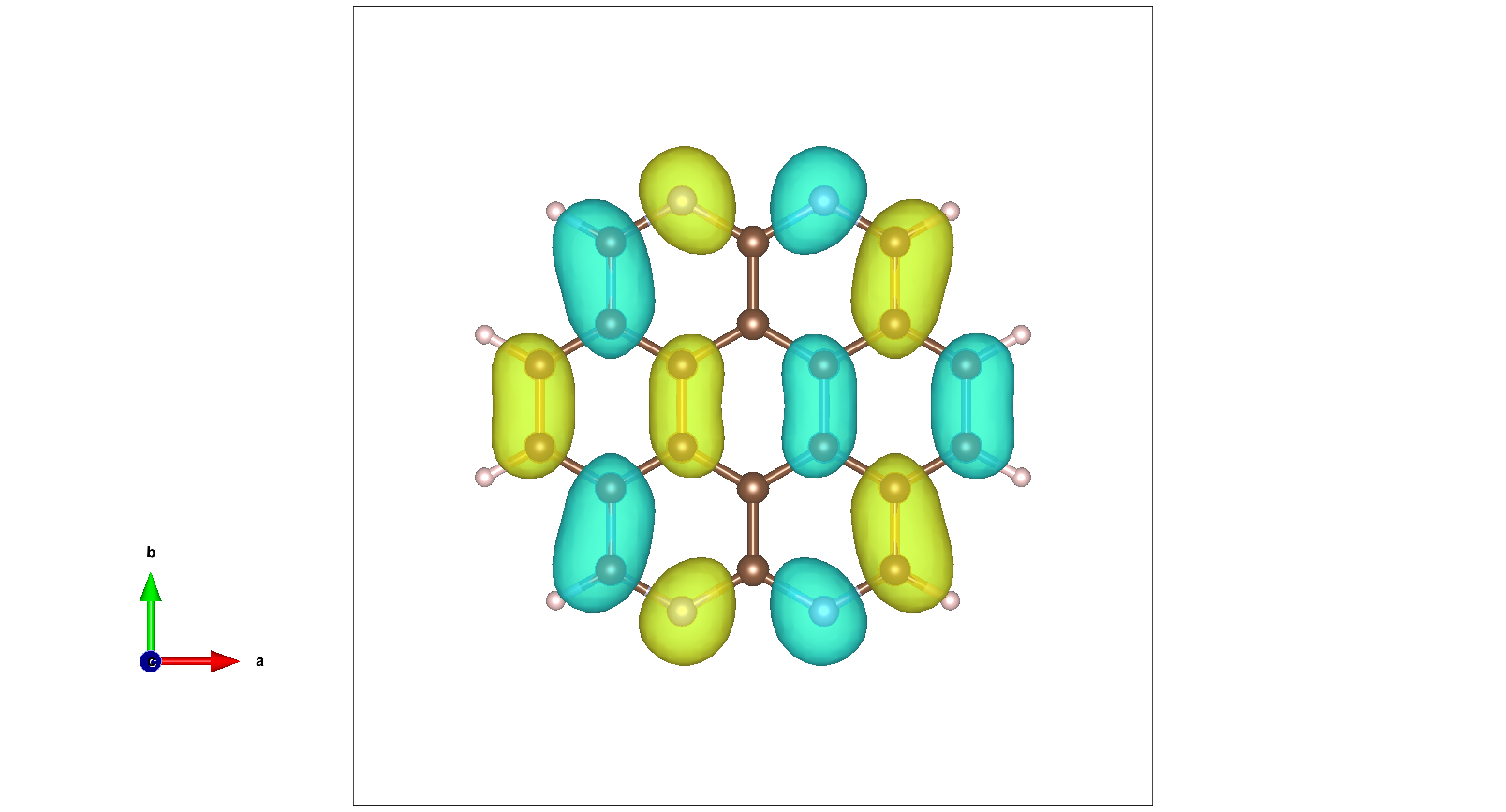
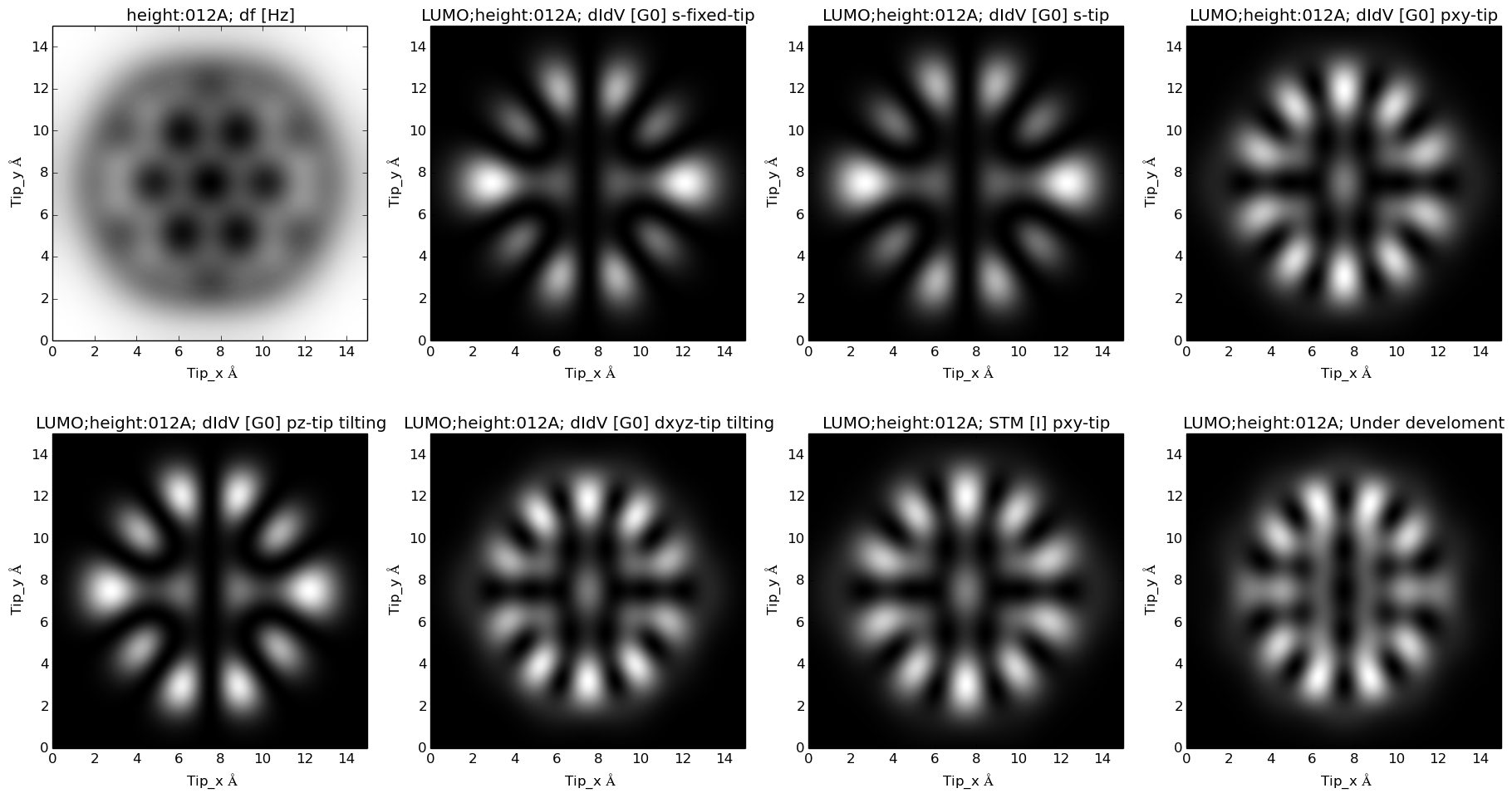
and LUMO: