Table of Contents
Constrain DFT
Constrained DFT allows to perform self consistent single electron excitation from arbitrary occupied orbital to unoccupied orbital (see example 1). There is also possibility to use quenching or geometry optimization of system in an excited state (see example 2).
Example 1
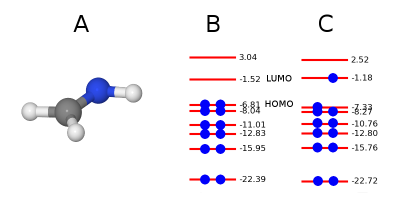
First, we show how to perform cDFT, i.e. SCF calculation of excited state, of formaldimine molecule CH2=NH. CH2=NH has in total 12 electron, from them 6 are double occupied molecular orbitals. Here we will run cDFT with a single excitation moving one electron from HOMO to LUMO state. The cDFT calculation is initiated by keywords icDFT = 1. See example of fireball.in file. Here is input file with initial geometry cnh3.bas.
fireball.in
&OPTION basisfile = CNH3.bas icluster = 1 nstepf = 1 T_initial = 100.0 T_final = 100.0 sigmatol = 0.000000001 dt = 0.01 iquench = 0 max_scf_iterations = 200 icDFT = 1 &END &OUTPUT iwrteigen = 1 iwrtxyz = 1 &END &MESH &END
To move electron from one orbital to another we need to create cDFT.optional input file in a directory. The first line means number of orbital where we want to create a state hole. The second line determines number of orbital ehwrw we want to create excited electron state. In this case there is missing one electron in HOMO orbital. The third line means occupancy excited electron.
6 ! hole state to be formed 7 ! excited electron state to be formed 1.0 ! occupancy of excited electron
Results of such calculation are on the fig[1]
Example 2
There is also possible to perform quenching of the excited state. The example is shown on the above mentioned system. CH2=N2 molecule can be found in two symetric conformations (Hydrogen bounded on N is in left or on the right side). Such transition is carried out by excitation of one electron from Homo to Lumo. Transition from one conformation to another pass through the conical intersection which is the local minimum of the excited state. In this example we will optimize geometry to the conical intersection. We only switch on keyword iquench = 1 and let them relax. Example of fireball in is below and elh.in file is the same as above:
&OPTION basisfile = CNH3.bas icluster = 1 nstepf = 5000 T_initial = 100.0 T_final = 100.0 sigmatol = 0.00000001 dt = 0.5 iquench = 1 max_scf_iterations = 200 iscfe = 1 &END &OUTPUT iwrteigen = 1 iwrtxyz = 1 &END &MESH &END